Genome Assembly of MRSA using Illumina MiSeq Data

Introduction to MRSA Genome Assembly using Illumina MiSeq Data
Introduction:
Methicillin-resistant Bacteria known as Staphylococcus aureus (MRSA) are resistant to the majority of antibiotics that are used to treat bacterial infections. Because of this, MRSA infections are challenging to treat and may result in serious complications. Understanding the molecular mechanisms underlying MRSA’s virulence, drug resistance, and evolution requires understanding its genome assembly. Due to its high throughput and accuracy, Illumina MiSeq is a well-liked platform for MRSA genome sequencing. Using data from Illumina MiSeq, we will introduce MRSA genome assembly in this article.
MRSA Genome Assembly:
Genome assembly is the process of piecing together fragments of DNA sequences to reconstruct the complete genome of an organism. The first step in MRSA genome assembly is to generate high-quality sequence data using Illumina MiSeq. The sequencing data generated by Illumina MiSeq is in the form of short reads, which are typically around 150-300 base pairs in length. The short reads need to be assembled into longer contiguous sequences, called contigs, which can be further scaffolded and ordered to generate a complete genome.
Data Pre-processing:
Before MRSA genome assembly, the raw sequencing data needs to be pre-processed to remove low-quality reads and adaptors. This is necessary to ensure that the assembly is of high quality and accuracy. The pre-processing steps include quality filtering, trimming, and error correction. Quality filtering removes low-quality reads based on parameters such as base quality scores, read length, and adapter content. Trimming removes adapter sequences and low-quality bases from the ends of reads. Error correction identifies and corrects sequencing errors to improve the accuracy of the assembly.
Genome Assembly Strategies:
De novo assembly and reference-based assembly are just two of the genome assembly techniques that can be applied to the MRSA genome using Illumina MiSeq data. The process of assembling the genome without using a reference genome is known as de novo assembly. This strategy is appropriate for novel or poorly understood organisms. In reference-based assembly, the sequencing reads are first aligned to a reference genome before being put together. For organisms with closely related reference genomes, this method is helpful.
Assembly Quality Control:
The quality of MRSA genome assembly using Illumina MiSeq data quality needs to be assessed to ensure that it is of high quality and accuracy. Several quality metrics can be used to evaluate the assembly, including contig length, genome coverage, and assembly accuracy. Contig length refers to the length of the contiguous sequence generated from the assembly. Genome coverage refers to the percentage of the genome that is covered by the assembly. Assembly accuracy refers to the correctness of the assembly with respect to the reference genome.
Final Analysis:
MRSA genome assembly using Illumina MiSeq data is an essential tool for understanding the molecular mechanisms underlying its virulence and drug resistance. The assembly process involves generating high-quality sequence data, pre-processing the data, assembling the genome, and evaluating the quality of the assembly. Genome assembly strategies and quality control measures need to be carefully selected to ensure that the assembly is of high quality and accuracy. MRSA genome assembly using Illumina MiSeq data has the potential to provide valuable insights into the biology and evolution of MRSA, which can be used to develop new strategies for preventing and treating MRSA infections.
Whole Genome Sequencing and MRSA Genome Assembly
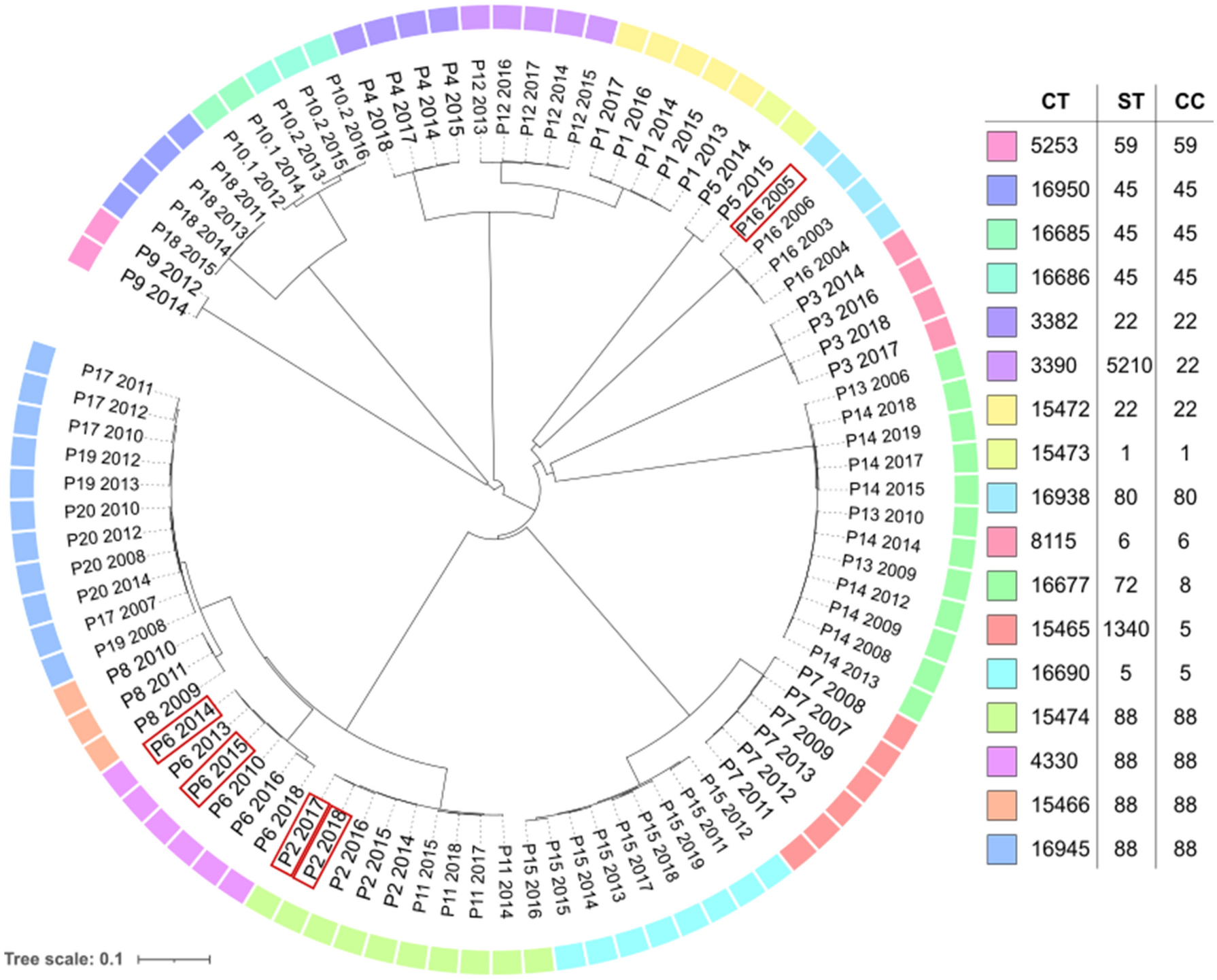
Whole genome sequencing (WGS) is a powerful tool used to study the genetic makeup of organisms. In the case of Methicillin-resistant Staphylococcus aureus (MRSA), a type of bacteria that is resistant to many antibiotics, WGS has been particularly useful for studying the entire genome of the bacteria in unprecedented detail. One important aspect of WGS in MRSA research is genome assembly, which involves reconstructing the entire genome sequence from a set of short DNA fragments.
To perform genome assembly, several steps are involved, including DNA extraction, library preparation, sequencing, and data analysis. During library preparation, the fragmented DNA is attached to adapters and loaded onto a flow cell for sequencing. The flow cell contains tiny channels where the DNA fragments are amplified and sequenced simultaneously, allowing for a large amount of data to be generated quickly and efficiently.
To achieve high-quality genome assemblies, it is essential to ensure that each DNA fragment is sequenced to the same sequence depth. This means that each fragment is sequenced the same number of times to ensure accurate and reliable assembly. Achieving the same sequence depth can be challenging, but it can be achieved by adjusting the amount of DNA loaded onto the flow cell.
After sequencing, the raw data is analyzed to identify and correct errors in the sequencing data. This involves comparing the sequence data to a reference database to improve accuracy and reduce the number of errors. Sequence similarity is an important factor in this analysis, as it helps identify and correct errors in the sequence data.
Finally, once the sequencing data has been analyzed, the genome can be assembled using specialized software and algorithms. This involves piecing together the short DNA fragments to reconstruct the entire genome sequence. In the case of MRSA, genome assembly can be particularly challenging due to the bacteria’s highly variable genome. However, by using specialized tools and techniques, high-quality genome assemblies can be achieved.
To sum it up, whole genome sequencing and MRSA genome assembly involve several important steps, including DNA extraction, library preparation, sequencing, and data analysis. Achieving the same sequence depth, analyzing sequence similarity, and using specialized tools and techniques are essential for generating high-quality genome assemblies. As the technology continues to advance, we can expect to see even more exciting developments in this field in the years to come.
Pre-processing of MRSA Sequence Data for Genome Assembly
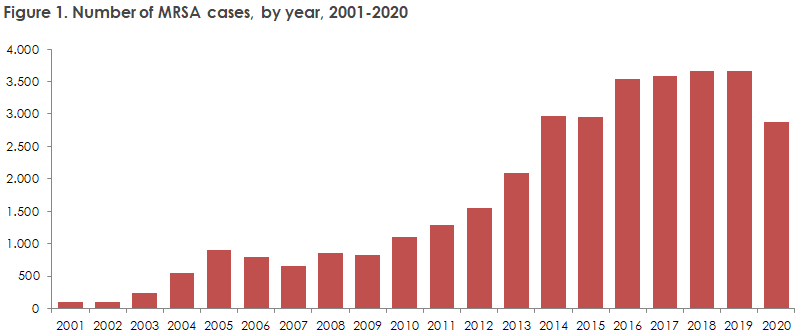
Pre-processing of MRSA sequence data is a critical step in the genome assembly process. This involves cleaning, filtering, and correcting errors in the raw sequence data to improve the accuracy and quality of the final genome assembly. In this article, we will explore the essential steps involved in pre-processing MRSA sequence data for genome assembly.
The first step in pre-processing MRSA sequence data is quality control. This involves assessing the quality of the raw sequence data to ensure that it meets the minimum standards required for accurate genome assembly. This is typically done using specialized software that evaluates the quality of the sequence reads based on several parameters, including read length, base quality, and sequencing depth.
Once the quality of the sequence data has been assessed, the next step is to filter out any low-quality reads or contaminants. This is essential to reduce the number of errors in the final genome assembly and improve the accuracy of the sequence data. This can be done using various filtering methods, such as removing reads with low base quality scores or those with low sequencing depth.
After filtering, the next step is to correct errors in the sequence data. This involves identifying and correcting errors in the base calls, such as mismatches or insertions, that can occur during sequencing. This can be done using specialized software that compares the sequence data to a reference genome or a set of high-quality reads to identify and correct errors in the sequence data.
Another critical step in pre-processing MRSA sequence data is to ensure that the reads are properly mapped to the reference genome. This involves aligning the sequence data to the reference genome to identify any structural variants or differences between the reference genome and the MRSA strain being sequenced. This can be done using specialized software that compares the sequence data to the reference genome and identifies any variants or differences.
Finally, the pre-processed sequence data is ready for genome assembly. This involves piecing together the sequence reads to reconstruct the entire genome sequence. The pre-processed sequence data is typically used as input for specialized software that uses various algorithms and tools to assemble the genome.
As a final point, pre-processing of MRSA sequence data is a critical step in the genome assembly process. Quality control, filtering, error correction, read mapping, and genome assembly are essential steps in pre-processing sequence data to ensure accurate and reliable genome assemblies. As the technology continues to advance, we can expect to see even more exciting developments in this field in the years to come.
Quality Control and Assessment of MRSA Sequencing Data quality assessment tool

The steps in the genome assembly process that involve quality control and assessment of MRSA sequencing data are essential. Researchers employ specialised quality assessment tools to guarantee the accuracy and dependability of the sequencing data generated. These tools help to increase the precision of the final genome assembly by enabling the detection of errors and inconsistencies in the sequencing data.
FastQC is one of the most frequently employed tools for evaluating the quality of MRSA sequencing data. By offering a variety of metrics, such as per-base sequence quality, per-base sequence content, adapter content, and sequence length distribution, this tool assesses the quality of the sequence reads. FastQC generates graphs and tables that allow researchers to spot any problems with the sequencing data, like adapter contamination or poor-quality reads.
Another popular quality assessment tool for MRSA sequencing data is Trimmomatic. This tool allows researchers to remove low-quality reads, adapters, and contaminants from the sequencing data. Trimmomatic uses a sliding window approach to identify regions with low-quality scores and removes them from the sequencing data. This improves the accuracy of the final genome assembly by eliminating reads that may introduce errors or inconsistencies.
In addition to these tools, researchers also use mapping and alignment tools, such as Bowtie2 and BWA, to assess the quality of the sequencing data. These tools allow for the identification of genomic variations, including SNPs and small insertions or deletions, which can be used to improve the accuracy of the final genome assembly.
Lastly, quality assurance and evaluation of MRSA sequencing data are critical phases in the genome assembly process. The accuracy of the final genome assembly can be determined by evaluating the quality of the sequencing data using specialized quality assessment tools like FastQC and Trimmomatic. The final genome assembly’s accuracy is increased by using mapping and alignment tools like Bowtie2 and BWA to find genomic variations. Researchers can obtain high-quality sequencing data by using these tools and the right protocols, which can then be used to analyze MRSA’s genetic makeup and create potent treatment plans.
Strategies for MRSA Genome Assembly using Illumina MiSeq Data
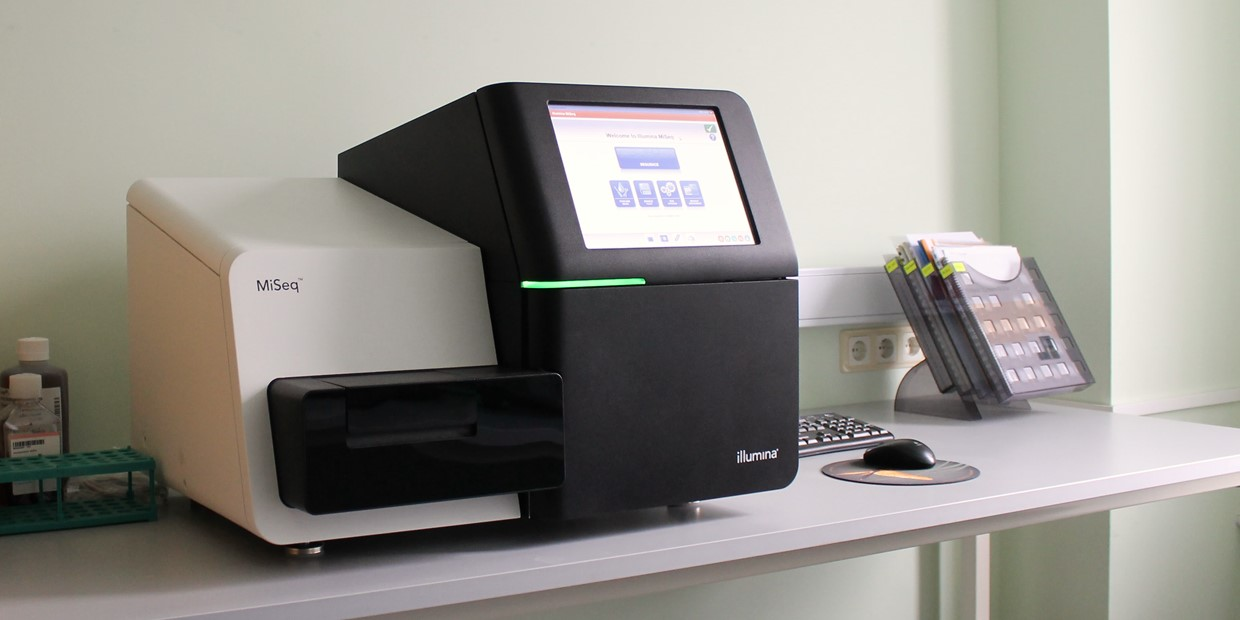

A difficult task that necessitates careful consideration of numerous factors is MRSA genome assembly using Illumina MiSeq data. A next-generation sequencing method called Illumina MiSeq can generate accurate and high-quality sequence data. The size of the MRSA genome, the existence of repetitive elements, and the substantial genetic diversity among MRSA strains, however, can make the genome assembly process challenging. We’ll talk about a few methods for assembling the MRSA genome using data from Illumina MiSeq in this article.
The first strategy is to optimize the sample preparation and sequencing protocol. This involves ensuring that the sequencing data produced has the same sequence depth across all regions of the genome. The sequencing depth should be sufficient to capture all variants present in the genome, including low-frequency variants. This can be achieved by optimizing the library preparation protocol and adjusting the sequencing parameters.
The second strategy is to perform quality control and assessment of the sequencing data. This involves evaluating the quality of the reads and identifying any issues that may affect the accuracy of the final genome assembly. Quality control and assessment can be performed using specialized software, such as FastQC and Trimmomatic, which can filter out low-quality reads and adapters.
The third tactic is to de novo assemble the MRSA genome using specialized software. Without a reference genome, the genome must be put together in this manner. De novo assembly of MRSA genomes can be accomplished using a number of software programmes, such as SPAdes and Velvet. To put together the genome and locate genomic variations, such as SNPs and small insertions or deletions, these software programmes employ a variety of algorithms.
The use of reference-guided assembly is the fourth tactic. In order to do this, the sequence reads must be aligned to a reference genome and genomic variations must be found. Large structural variations in the genome can be found and repetitive regions can be resolved using this method.
MRSA genome assembly using Illumina MiSeq data can be challenging, but several strategies can be used to improve the accuracy and reliability of the final genome assembly. By optimizing the sample preparation and sequencing protocol, performing quality control and assessment of the sequencing data, using specialized software for de novo assembly or reference-guided assembly, researchers can obtain high-quality sequence data that can be used to study the genetics of MRSA and develop effective treatment strategies.
Optimizing Sequence Data Analysis for MRSA Genome Assembly
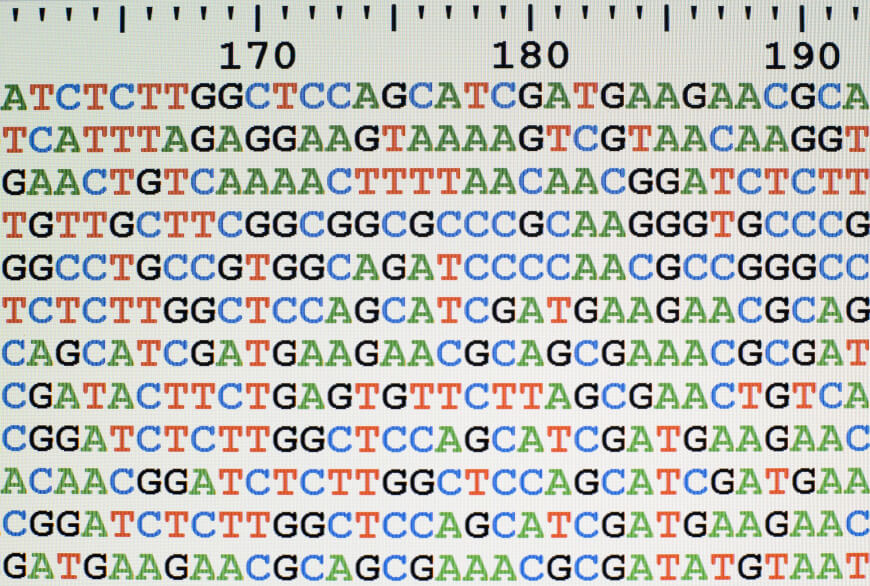
The analysis of sequence data must be optimized before the MRSA genome can be assembled. The reliability of the sequence data and the efficiency of the data analysis tools used to determine how accurate and comprehensive the genome assembly is. We will go over a few methods for streamlining sequence data analysis for MRSA genome assembly in this article.
The sequencing data must first be subjected to quality control and evaluation. This entails assessing the read quality and looking for any problems that might compromise the accuracy of the final genome assembly. Specialized software that can weed out low-quality reads and adapters, like Trimmomatic and FastQC, can be used for quality control and evaluation.
The second step is to select an appropriate genome assembly tool. Several software packages are available for genome assembly, including de novo assembly and reference-guided assembly. The choice of assembly tool depends on the sequencing data, the size of the genome, and the availability of a reference genome.
The third step is to perform error correction of the sequencing data. This involves correcting any errors in the sequence data that may affect the accuracy of the genome assembly. Error correction can be performed using specialized software, such as BayesHammer and SOAPec, which can correct errors in the sequencing data by comparing the reads to each other.
The fourth step is to identify and resolve genomic variations, including single nucleotide polymorphisms (SNPs), insertions, and deletions. Several software packages, such as GATK and Samtools, can be used to identify genomic variations and perform variant calling.
The fifth step is to annotate the genome. Genome annotation involves identifying and labeling the genes and other functional elements in the genome. Several software packages, such as Prokka and RAST, can be used for genome annotation.
It is essential for MRSA genome assembly to optimise sequence data analysis. Researchers can obtain high-quality sequence data that can be used to study the genetics of MRSA and create efficient treatment plans by performing quality control and assessment of the sequencing data, choosing an appropriate assembly tool, performing error correction, identifying and resolving genomic variations, and annotating the genome.
During the analysis of MRSA sequencing data, it is important to carefully evaluate the results for statistically significant differences, as well as to take into account any other third-party material that may impact the accuracy of the data. Next-generation sequencing technologies have revolutionized our ability to study the genetics of MRSA, but the accuracy and reliability of the data can be influenced by a variety of factors, including sample quality and experimental design. To ensure that the results are accurate and reliable, it is essential to carefully control for these factors and to use rigorous statistical analysis techniques to evaluate the data.
Additionally, researchers must be aware of the potential impact of other third-party material, such as contaminants or environmental factors, on the sequencing data, and take steps to minimize their impact on the analysis. By carefully controlling for these factors and using rigorous analysis techniques, researchers can obtain high-quality data that can be used to better understand the genetics of MRSA and develop new treatment strategies.
Evaluation of MRSA Genome Assembly Quality using Illumina MiSeq Data
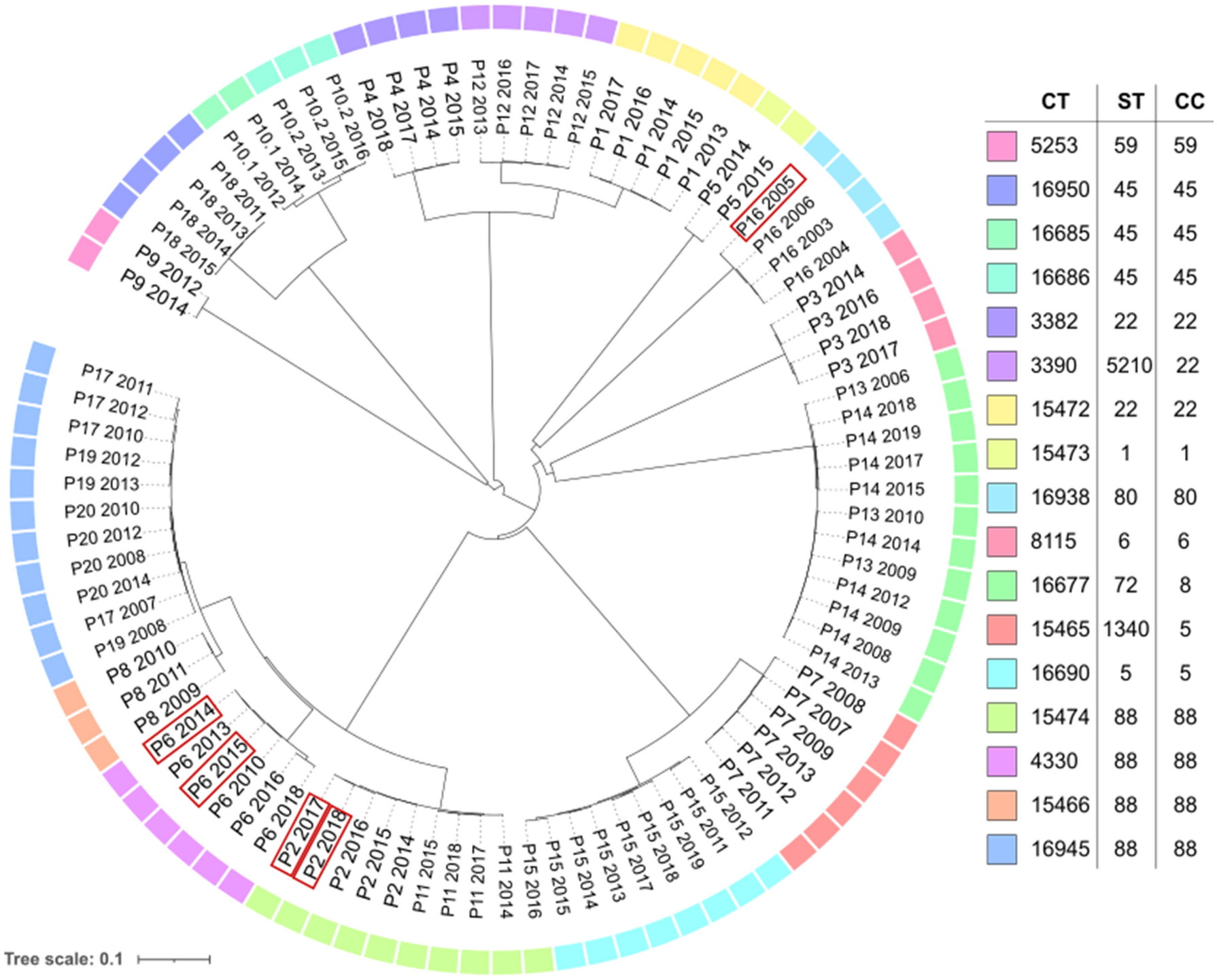
Understanding the pathogenesis and evolution of MRSA strains requires the use of an important genetic research technique called MRSA genome assembly. High-throughput sequencing of MRSA genomes has increased in popularity with the introduction of next-generation sequencing technologies like Illumina MiSeq. However, a number of variables, including read length, sequencing depth, and the calibre of the input data, can affect the quality of the genome assembly. To ensure the accuracy and completeness of the data, it is crucial to assess the quality of the MRSA genome assembly.
One way to evaluate the quality of the MRSA genome assembly is to assess the assembly metrics, such as the contig N50 and the number of contigs. The N50 is a metric that represents the length of the shortest contig needed to cover 50% of the genome assembly. The number of contigs is an indicator of the completeness of the genome assembly. High-quality assemblies typically have a high N50 and a low number of contigs.
Comparing the MRSA genome assembly to a reference genome is another way to assess its quality. This can be accomplished by comparing the assembled genome to the reference genome using a variety of software tools, including Mauve and QUAST. Researchers can find potential assembly errors and fix them to increase the accuracy of the data by comparing the assembled genome to a reference genome.
Researchers can also assess the quality of the genome assembly visually with the aid of visualisation tools like Artemis. Visualization tools can be used to spot potential assembly mistakes and manually fix any that are made.
So, the quality of the MRSA genome assembly is critical for accurate and reliable genetic analysis. By evaluating the assembly metrics, comparing the assembly to a reference genome, and using visualization tools, researchers can ensure the accuracy and completeness of the data. This will allow for a better understanding of the genetics of MRSA and the development of effective treatment strategies.
Annotation and Functional Analysis of MRSA Genome using Sequencing Data

An essential step in genetic research is the annotation and functional analysis of the MRSA genome, which can reveal important information about the pathogenesis and evolution of MRSA strains. High-throughput sequencing of MRSA genomes has increased in popularity since the development of next-generation sequencing technologies, giving researchers access to enormous amounts of genetic data that can be used for annotation and functional analysis.
Genes, regulatory regions, and non-coding regions are just a few examples of the various features of the genome that are identified and labelled during annotation. The location and function of each gene can be predicted using software tools like Prokka and RAST, which use algorithms based on sequence similarity and other features. Annotation provides a foundation for further analysis and can help researchers to identify potential virulence factors and drug resistance genes.
Functional analysis involves interpreting the annotation data to understand the biological functions of the genes and their interactions with other genes and proteins. This can be done using various bioinformatics tools, such as KEGG and COG, which can identify functional pathways and networks within the genome. Functional analysis can help researchers to understand the molecular mechanisms underlying virulence and drug resistance, and to identify potential targets for new treatments.
In addition, the use of comparative genomics, which involves comparing the MRSA genome to other closely related genomes, can provide additional insights into the functional significance of genetic variations. Comparative genomics can help to identify conserved and variable regions within the genome, as well as potential horizontal gene transfer events.
An important step in genetic research that can yield important insights into the pathogenesis and evolution of MRSA strains is the annotation and functional analysis of the MRSA genome using sequencing data. Researchers can create fresh treatments and tactics to fight MRSA infections by locating potential virulence factors and drug resistance genes, comprehending functional pathways and networks, and using comparative genomics to find genetic variations.
Comparative Genomics of MRSA Strains using Whole Genome Sequencing
Comparative genomics is an important approach to understanding the genetic diversity and evolution of MRSA strains. With the availability of whole genome sequencing data, researchers can now compare the genetic makeup of multiple MRSA strains to identify similarities and differences in their genomes.
Comparative genomics can provide insights into the evolution of MRSA strains, including the acquisition of resistance genes and the development of virulence. By comparing the genomes of different MRSA strains, researchers can identify common genetic markers and pathways that may be associated with virulence or resistance, as well as unique features that may be specific to certain strains.
In addition, comparative genomics can help to identify potential sources of MRSA transmission, including hospital outbreaks and community-acquired infections. By comparing the genomes of MRSA strains from different locations and time periods, researchers can identify patterns of transmission and potentially trace the origins of new outbreaks.
Furthermore, comparative genomics can be used to develop new diagnostic tools and therapies for MRSA infections. By identifying specific genetic markers associated with virulence or resistance, researchers can develop targeted diagnostic tests and therapies that are more effective at treating MRSA infections.
Overall, comparative genomics is a powerful tool for understanding the genetic diversity and evolution of MRSA strains. By analyzing the genomes of multiple MRSA strains, researchers can identify common features and unique variations that provide insights into the pathogenesis and transmission of MRSA, as well as new opportunities for diagnosis and treatment.
Insights into MRSA Pathogenicity through Genome Assembly and Data Analysis
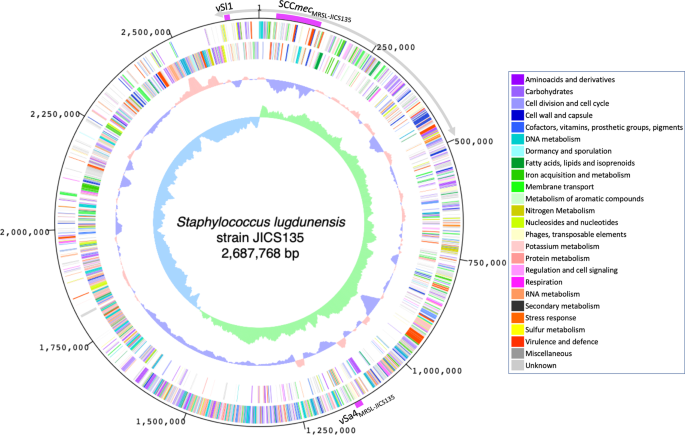
Methicillin-resistant Staphylococcus aureus (MRSA) is a highly virulent bacterium that poses a significant threat to public health. To better understand the pathogenicity of MRSA and develop new treatments, researchers are using genome assembly and data analysis to gain insights into the genetic makeup of MRSA strains.
Genome assembly involves piecing together the individual nucleotide sequences of an organism’s genome to create a complete picture of its genetic makeup. With next-generation sequencing technologies, researchers can sequence the entire MRSA genome in a single experiment. This provides a vast amount of genetic data that can be used to identify potential virulence factors and drug resistance genes.
Data analysis is then used to interpret the genetic data and identify potential targets for new treatments. This includes identifying genetic variations between different MRSA strains and understanding how these variations affect the bacterium’s virulence and resistance. By understanding the molecular mechanisms underlying MRSA pathogenicity, researchers can develop new drugs and therapies that are more effective at treating MRSA infections.
Furthermore, genome assembly and data analysis can be used to track the transmission of MRSA infections, which is critical for identifying and controlling outbreaks. By comparing the genomes of MRSA strains from different locations and time periods, researchers can identify patterns of transmission and trace the origins of new outbreaks.
Overall, genome assembly and data analysis are powerful tools for gaining insights into MRSA pathogenicity and developing new treatments. By identifying potential virulence factors and drug resistance genes, understanding genetic variations, and tracking the transmission of MRSA infections, researchers can make significant progress towards combating this dangerous bacterium .
Future Directions for MRSA Genome Assembly and Sequencing Data Analysis
In the field of MRSA genome assembly and sequencing data analysis, there are several exciting future directions that hold promise for advancing our understanding of this dangerous bacterium. One key area of development is the continued improvement of next-generation sequencing technologies, with a focus on generating longer reads and higher sequencing depths to capture more complete information about the MRSA genome.
Another important direction is the development of new annotation and functional analysis tools to better interpret the vast amount of sequencing data generated by MRSA experiments. This includes the development of new algorithms to identify potential virulence factors and drug resistance genes, as well as tools for predicting the functional consequences of genetic variations.
In addition, the integration of multi-omics data, such as transcriptomics and proteomics, is an exciting area of development that can provide a more comprehensive understanding of MRSA pathogenicity. By combining different types of data, researchers can better identify key molecular pathways and potential drug targets.
Ultimately, continued collaboration and data sharing between researchers will be key to making further progress in the field of MRSA genome assembly and sequencing data analysis. With these exciting developments on the horizon, we can look forward to a brighter future in the fight against MRSA and other deadly bacterial pathogens.
Conclusion and Implications of MRSA Genome Assembly using Illumina MiSeq Data and Data Analysis.

Illumina MiSeq sequencing technology, developed in San Diego CA USA, has revolutionized the field of targeted sequencing, allowing for efficient and accurate analysis of specific genomic regions in MRSA strains. This powerful tool enables researchers to quickly identify genetic variants associated with drug resistance and virulence, providing important insights into the mechanisms underlying MRSA pathogenesis. By leveraging the power of Illumina MiSeq sequencing, researchers in San Diego and around the world can continue to make significant strides in our understanding of MRSA and develop new strategies for controlling and preventing infections.
Using Illumina MiSeq data to assemble and analyse the MRSA genome has great potential to advance our knowledge of this pathogen. This sequencing technology enables the identification of important genomic features linked to virulence and drug resistance due to its high throughput and accuracy.
We can learn more about the genetic components influencing MRSA’s pathogenicity through data analysis, which will help us develop new diagnostic techniques and therapeutic plans. Additionally, comparative genomics can assist in identifying distinctive genomic traits linked to particular MRSA strains, supplying crucial information for surveillance and outbreak management.
However, it is important to acknowledge that there are still challenges to be addressed in MRSA genome assembly and data analysis, such as the need for more efficient and accurate annotation tools and the integration of multi-omics data. Additionally, careful consideration must be given to the ethical implications of using genomic data, particularly in terms of patient privacy and data sharing.
In the end, the use of Illumina MiSeq data and data analysis provides a valuable tool for advancing our understanding of MRSA and improving patient outcomes. By continuing to refine and improve this technology, we can hope to make significant progress in the fight against this deadly pathogen.